LymphAware: Domain-Aware Bias Disruption for Reliable Lymphoma Cancer AI Diagnosis
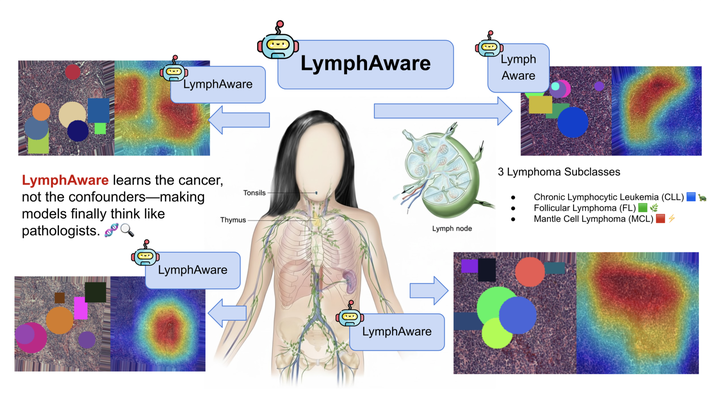
Abstract
Lymphoma histopathology classifiers frequently achieve high accuracy by exploiting non-biological visual cues such as scanner color signatures, stain-saturation drift, background texture, or slide-processing artifacts. These shortcut signals inflate in-domain performance while causing brittle behavior under cross-acquisition domain shift. We propose LymphAware, a domain-aware bias disruption framework that explicitly separates lymphoma-relevant morphology from shortcut-sensitive acquisition factors. The method is designed for realistic multi-source histopathology settings, without assuming verified institutional separation. LymphAware integrates three complementary mechanisms, (1) morphology-centric feature isolation, (2) adversarial and orthogonality-based shortcut suppression, and (3) cross-domain stability constraints that enforce acquisition-invariant representations. Each component addresses a distinct shortcut failure mode, avoiding redundant architectural complexity. To expose latent shortcut dependence, we introduce artifact-shift perturbations that simulate staining and scanner variability and enforce counterfactual consistency during training. Evaluated on a heterogeneous lymphoma histopathology benchmark, LymphAware consistently improves cross-domain robustness, degrades gracefully under loss-weight variation, and yields attribution maps aligned with pathology-relevant regions. We emphasize that these attributions reflect associative alignment rather than proven causal inference. These results highlight the importance of representation-level shortcut disruption for reliable and clinically meaningful lymphoma diagnosis.


You can explore the full LymphAware project at 👉 https://kaopanboonyuen.github.io/LymphAware/
The official implementation is publicly available at 👉 https://github.com/kaopanboonyuen/LymphAware
Unlike conventional histopathology classifiers that implicitly rely on acquisition-specific shortcuts, LymphAware is designed from the ground up to disentangle morphology from scanner- and stain-induced bias. The repository includes reproducible training pipelines, artifact-shift perturbation modules, and evaluation protocols for cross-domain robustness. Our goal is not merely higher accuracy, but representation-level reliability under realistic domain shift — a necessary step toward clinically trustworthy AI systems.